Association of Polymorphisms in the 3' UTR of Genes in the ERK1/2 Signaling Pathway with Non-small Cell Lung Cancer
-
摘要:
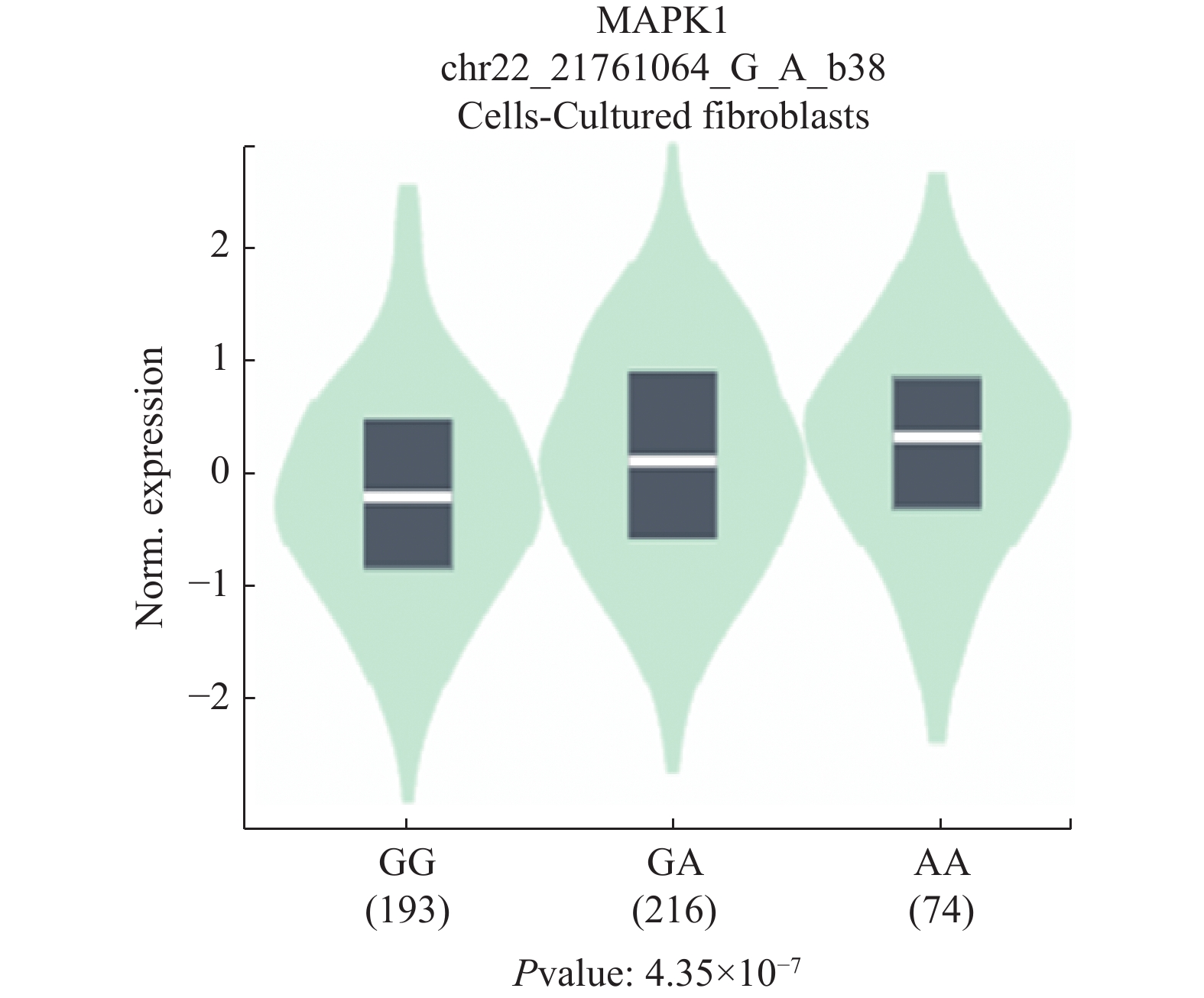
目的 探究4个ERK1/2信号通路的基因3'UTR区域的单核苷酸多态性(single nucleotide polymorphism,SNP)位点(MAPK1基因中的rs9340,NRAS基因中的rs14804,KRAS基因中的rs712和rs7973450)与非小细胞肺癌(non-small cell lung cancer,NSCLC)的相关性。 方法 纳入了478名NSCLC患者及480名健康对照者,利用TaqMan探针法对其进行基因分型检测,并分析上述4个SNP与NSCLC的相关性。 结果 rs9340位点的等位基因在对照组与非小细胞鳞状细胞癌组( squamous cell carcinoma,SCC)中分布频率的差异具有统计学意义(P = 0.009),该结果表明rs9340 位点的G等位基因可能是非小细胞肺鳞癌的保护性因素(OR = 0.67,95% CI: 0.50~0.91)。同时,在 < 50岁年龄组中,rs9340位点的等位基因在对照组和NSCLC组中的分布频率差异具有统计学意义(P = 5.07 × 10-4),该结果表明rs9340等位基因G可能是NSCLC的保护性因素(OR = 0.46,95% CI: 0.29~0.72)。 结论 MAPK1基因SNP位点rs9340可能与NSCLC的发生风险相关。 -
关键词:
- 非小细胞肺癌 /
- ERK1/2信号通路 /
- 单核苷酸多态性 /
- 3'UTR /
- MAPK1
Abstract:Objective To investigate the association between four single nucleotide polymorphisms(SNP)(rs9340 in MAPK1, rs14804 in NRAS, rs712 and rs7973450 in KRAS) in the 3'UTR of ERK1/2 signaling pathway-related genes and non-small cell lung cancer(NSCLC). Methods A total of 478 NSCLC patients and 480 healthy controls were enrolled in this study. Four SNPs were genotyped by using TaqMan assays. The association between the four SNPs and NSCLC was analyzed. Results The distribution frequency difference of the allele of rs9340 was statistically significant between the control group and the non-small cell squamous cell carcinoma(SCC) group(P = 0.009), suggesting that the G allele of rs9340 may be a protective factor for non-small cell lung squamous cell carcinoma(OR = 0.67, 95%CI: 0.50~0.91). In addition, in the < 50 years age group, the distribution frequency difference of the allele of rs9340 was statistically significant between the control group and the NSCLC group(P = 5.07 × 10-4), indicating that the G allele of rs9340 may be a protective factor for NSCLC(OR = 0.46, 95%CI: 0.29~0.72). Conclusion The SNP rs9340 in MAPK1 may be associated with the risk of NSCLC. -
Key words:
- Non-small cell lung cancer /
- ERK1/2 signaling pathway /
- SNP /
- 3'UTR /
- MAPK1
-
表 1 所选SNP位点信息
Table 1. The information of selected SNPs in the current study
SNPs 基因 功能 位置 等位基因 中国南方汉族人群MAF rs9340 MAPK1 3'UTR突变 Chr 22:21761064 G > A 0.21 rs14804 NRAS 3'UTR突变 Chr 1:114707222 G > A 0.04 rs712 KRAS 3'UTR突变 Chr 12:25209618 A > C 0.48 rs7973450 KRAS 3'UTR突变 Chr 12:25208208 A > G 0.09 SNP,单核苷酸多态性;MAF,次要等位基因频率;Chr,染色体。 表 2 受试者临床信息[$ x \pm s $/n(%)]
Table 2. The clinical characteristics of the subjects enrolled in the current study [$ x \pm s $/n(%)]
项目 肺癌组 对照组 t/χ2 P 样本数 478 480 年龄(岁) 55.94 ± 10.79 55.16 ± 11.35 1.09 0.275 性别 男 305(63.8) 277(57.7) 3.74 0.053 女 173(36.2) 203(42.3) 病理类型 腺癌 327(68.4) 8.00 < 0.001* 鳞状细胞癌 151(31.6) 临床分期 Ⅰ期 126(26.3) 26.45 < 0.001* Ⅱ期 74(15.5) Ⅲ期 151(31.6) Ⅳ期 127(26.6) 表 3 在健康对照组和NSCLC组中4个SNP位点等位基因及基因型频率分布结果[n(%)]
Table 3. Allele and genotype frequencies of four SNPs between the control group and NSCLC group [n(%)]
SNPs 等位基因/基因型 对照组 肺癌组 对照组HWE 肺癌组 vs 对照组 χ2 P OR (95%CI) χ2 P rs9340 G 773(80.5) 728(76.2) 0.77(0.62~0.96) 5.39 0.020 A 187(19.5) 228(23.8) G/G 308(64.2) 273(57.1) 0.87 0.350 5.63 0.060 G/A 157(32.7) 182(38.1) A/A 15(3.1) 23(4.8) rs14804 G 936(97.5) 935(97.8) 1.14(0.63~2.06) 0.19 0.661 A 24(2.5) 21(2.2) G/G 456(95.0) 458(95.8) 0.32 0.574 1.58 0.454 G/A 24(5.0) 19(4.0) A/A 0 1(0.2) rs712 A 197(20.5) 202(21.1) 1.04(0.83~1.29) 1.04 0.743 C 763(79.5) 754(78.9) A/A 21(4.4) 29(6.1) 0.05 0.826 1.68 0.431 A/C 155(32.3) 144(30.1) C/C 304(63.3) 305(63.8) rs7973450 A 873(90.9) 876(91.6) 1.09(0.79~1.50) 1.09 0.590 G 87(9.1) 80(8.4) A/A 399(83.1) 401(83.9) 1.30 0.254 1.01 0.604 A/G 75(15.6) 74(15.5) G/G 6(1.3) 3(0.6) HWE,哈迪温伯格平衡。*P < 0.012(经Bonferroni校正,n = 4)。 表 4 在健康对照组和不同病理类型肺癌组中4个SNP位点等位基因及基因型频率分布结果[n(%)]
Table 4. Allele and genotype frequencies of four SNPs between different pathological types of NSCLC [n(%)]
SNPs 等位基因/基因型 对照组 AC组 SCC组 AC组 vs 对照组 SCC组 vs 对照组 SCC组 vs AC组 OR(95%CI) χ2 P OR(95%CI) χ2 P OR(95%CI) χ2 P rs9340 G 773(80.5) 506(77.4) 222(73.5) 0.83(0.65~1.05) 2.35 0.125 0.67(0.50~0.91) 6.77 0.009* 0.81(0.59~1.11) 1.70 0.193 A 187(19.5) 148(22.6) 80(26.5) G/G 308(64.2) 192(58.7) 81(53.7) 2.53 0.282 7.26 0.027 2.13 0.345 G/A 157(32.7) 122(37.3) 60(39.7) A/A 15(3.1) 13(4.0) 10(6.6) rs14804 G 936(97.5) 640(97.9) 295(97.7) 1.17(0.60~2.28) 0.22 0.640 1.08(0.46~2.53) 0.03 0.858 0.92(0.37~2.31) 0.03 0.862 A 24(2.5) 14(2.1) 7(2.3) G/G 456(95.0) 313(95.7) 145(96.0) 0.22 0.636 3.90 0.142 2.41 0.300 G/A 24(5.0) 14(4.3) 5(3.3) A/A 0 0 1(0.7) rs712 A 197(20.5) 140(21.4) 62(20.5) 1.05(0.83~1.35) 0.18 0.667 1.00(0.73~1.38) 1.10 × 10−5 0.997 0.95(0.68~1.33) 0.09 0.758 C 763(79.5) 514(78.6) 240(79.5) A/A 21(4.4) 20(6.1) 9(6.0) 1.34 0.512 1.01 0.602 0.12 0.943 A/C 155(32.3) 100(30.6) 44(29.1) C/C 304(63.3) 207(63.3) 98(64.9) rs7973450 A 873(90.9) 597(91.3) 279(92.4) 1.04(0.74~1.48) 0.06 0.810 1.21(0.75~1.95) 0.60 0.437 1.16(0.70~1.92) 0.32 0.568 G 87(9.1) 57(8.7) 23(7.6) A/A 399(83.1) 273(83.5) 128(84.8) 0.20 0.907 1.94 0.380 1.41 0.493 A/G 75(15.6) 51(15.6) 23(15.2) G/G 6(1.3) 3(0.9) 0 AC,肺腺癌;SCC,肺鳞状细胞癌。*P < 0.012(经Bonferroni校正,n = 4)。 表 5 健康对照组和不同分期肺癌组中4个SNP位点等位基因及基因型频率分布结果[n(%)]
Table 5. Allele and genotype frequencies of four SNPs between different stages of NSCLC [n(%)]
SNPs 等位基因/基因型 对照组 I+II期组 III+IV期组 I+II期组 vs 对照组 III+IV期组 vs 对照组 III+IV期组 vs I+II期组 OR(95%CI) χ2 P OR(95%CI) χ2 P OR(95%CI) χ2 P rs9340 G 773(80.5) 303(75.7) 425(76.4) 0.76(0.57~1.00) 3.89 0.049 0.78(0.61~1.01) 3.54 0.060 1.04(0.77~1.40) 0.06 0.805 A 187(19.5) 97(24.3) 131(23.6) G/G 308(64.2) 114(57.0) 159(57.2) 4.20 0.122 3.78 0.151 0.37 0.831 G/A 157(32.7) 75(37.5) 107(38.5) A/A 15(3.1) 11(5.5) 12(4.3) rs14804 G 936(97.5) 394(98.5) 541(97.3) 1.68(0.68~4.15) 1.31 0.253 0.92(0.48~1.78) 0.05 0.815 0.55(0.21~1.43) 1.55 0.213 A 24(2.5) 6(1.5) 15(2.7) G/G 456(95.0) 194(97.0) 264(95.0) 1.34 0.247 1.76 0.414 1.59 0.451 G/A 24(5.0) 6(3.0) 13(4.6) A/A 0 0 1(0.4) rs712 A 197(20.5) 79(19.8) 123(22.1) 0.95(0.71~1.28) 0.10 0.747 1.10(0.85~1.42) 0.54 0.462 1.15(0.84~1.59) 0.79 0.375 C 763(79.5) 321(80.2) 433(77.9) A/A 21(4.4) 9(4.5) 20(7.2) 0.21 0.901 2.92 0.232 1.48 0.476 A/C 155(32.3) 61(30.5) 83(29.9) C/C 304(63.3) 130(65.0) 175(62.9) rs7973450 A 873(90.9) 370(92.5) 506(91.0) 1.23(0.80~1.89) 0.88 0.349 1.01(0.70~1.45) 2.08 × 10−3 0.964 0.82(0.51~1.32) 0.68 0.411 G 87(9.1) 30(7.5) 50(9.0) A/A 399(83.1) 173(86.5) 228(82.0) 1.53 0.465 4.10 0.129 7.14 0.028 A/G 75(15.6) 24(12.0) 50(18.0) G/G 6(1.3) 3(1.5) 0 *P < 0.012(经Bonferroni校正,n = 4)。 表 6 不同年龄组中4个SNP位点等位基因及基因型频率分布结果[n(%)] (1)
Table 6. Allele and genotype frequencies of four SNPs between different age groups [n(%)] (1)
SNPs 等位基因/
基因型< 50岁组 50~65岁组 > 65岁组 < 50岁
对照组vs肺癌组50~65岁
对照组vs肺癌组> 65岁
对照组vs肺癌组对照组 肺癌组 对照组 肺癌组 对照组 肺癌组 OR(95%CI) χ2 P OR(95%CI) χ2 P OR(95%CI) χ2 P rs9340 G 220(85.9) 196
(73.7)387
(79.3)380
(75.4)166(76.9) 152(81.7) 0.46(0.29~0.72) 12.10 5.07 × 10−4* 0.80(0.59~1.08) 2.16 0.142 1.35
(0.83~2.19)1.43 0.231 A 36(14.1) 70
(26.3)101
(20.7)124
(24.6)50
(23.1)34
(18.3)G/G 92(71.9) 70
(52.6)151
(61.9)144
(57.2)65
(60.2)59
(63.4)14.24 8.15 × 10−4 2.98 0.225 6.26 0.044 G/A 36(28.1) 56
(42.1)85
(34.8)92
(36.5)36
(33.3)34
(36.6)A/A 0 7
(5.3)8
(3.3)16
(6.3)7
(6.5)0 rs14804 G 248(96.9) 260
(97.7)475
(97.3)490
(97.2)213
(98.6)185
(99.5)1.40
(0.48~4.09)0.38 0.539 0.96
(0.45~2.06)0.01 0.912 2.61
(0.27~25.27)0.74 0.391 A 8(3.1) 6
(2.3)13
(2.7)14
(2.8)3
(1.4)1
(0.5)G/G 120(93.8) 127
(95.5)231
(94.7)239
(94.8)105
(97.2)92
(98.9)0.39 0.533 1.04 0.592 0.74 0.389 G/A 8(6.2) 6
(4.5)13
(5.3)12
(4.8)3
(2.8)1
(1.1)A/A 0 0 0 1
(0.4)0
(0.0)0 表 6 不同年龄组中4个SNP位点等位基因及基因型频率分布结果[n(%)] (2)
Table 6. Allele and genotype frequencies of four SNPs between different age groups [n(%)] (2)
SNPs 等位基因/
基因型< 50岁组 50~65岁组 > 65岁组 < 50岁
对照组vs肺癌组50~65岁
对照组vs肺癌组> 65岁
对照组vs肺癌组对照组 肺癌组 对照组 肺癌组 对照组 肺癌组 OR(95%CI) χ2 P OR(95%CI) χ2 P OR(95%CI) χ2 P rs712 A 53(20.7) 50
(18.8)95
(19.5)111
(22.0)49
(22.7)41
(22.0)0.89
(0.58~1.36)0.30 0.584 1.17
(0.86~1.59)0.98 0.321 0.96
(0.60~1.54)0.02 0.878 C 203(79.3) 216
(81.2)393
(80.5)393
(78.0)167
(77.3)145
(78.0)A/A 8(6.3) 4
(3.0)6
(2.5)19
(7.5)7
(6.5)6
(6.4)1.65 0.439 7.35 0.025 0.04 0.982 A/C 37(28.9) 42
(31.6)83
(34.0)73
(29.0)35
(32.4)29
(31.2)C/C 83(64.8) 87
(65.4)155
(63.5)160
(63.5)66
(61.1)58
(62.4)rs7973450 A 233(91.0) 246
(92.5)441
(90.4)459
(91.1)199
(92.1)171
(91.9)1.21
(0.65~2.27)0.37 0.543 1.09
(0.71~1.67)0.15 0.703 0.97
(0.47~2.01)5.14 × 10−3 0.943 G 23(9.0) 20
(7.5)47
(9.6)45
(8.9)17
(7.9)15
(8.1)A/A 108(84.4) 114
(85.7)198
(81.2)209
(82.9)93
(86.1)78
(83.9)1.10 0.578 0.69 0.709 2.35 0.309 A/G 17(13.3) 18
(13.5)45
(18.4)41
(16.3)13
(12.0)15
(16.1)G/G 3(2.3) 1
(0.8)1
(0.4)2
(0.8)2
(1.9)0 *P < 0.012(经Bonferroni校正,n = 4)。 -
[1] Sung H,Ferlay J,Siegel R L,et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin,2021,71(3):209-249. doi: 10.3322/caac.21660 [2] Zheng R,Zhang S,Zeng H,et al. Cancer incidence and mortality in China,2016[J]. Journal of the National Cancer Center,2022,2(1):1-9. doi: 10.1016/j.jncc.2022.02.002 [3] Nooreldeen R,Bach H. Current and future development in lung cancer diagnosis[J]. Int J Mol Sci,2021,22(16):8661. doi: 10.3390/ijms22168661 [4] Cao W,Chen H D,Yu Y W,et al. Changing profiles of cancer burden worldwide and in China: A secondary analysis of the global cancer statistics 2020[J]. Chin Med J (Engl),2021,134(7):783-791. doi: 10.1097/CM9.0000000000001474 [5] Saller J J,Boyle T A. Molecular pathology of lung cancer[J]. Cold Spring Harb Perspect Med,2022,12(3):a037812. doi: 10.1101/cshperspect.a037812 [6] Duma N,Santana-Davila R,Molina JR. Non-small cell lung cancer: Epidemiology,screening,diagnosis,and treatment[J]. Mayo Clin Proc,2019,94(8):1623-1640. doi: 10.1016/j.mayocp.2019.01.013 [7] Zhang J,Chen S F,Zhen Y,et al. Multicenter analysis of lung cancer patients younger than 45 years in Shanghai[J]. Cancer,2010,116(15):3656-3662. doi: 10.1002/cncr.25100 [8] Heist R S,Sequist L V,Engelman J A. Genetic changes in squamous cell lung cancer: a review[J]. J Thorac Oncol,2012,7(5):924-933. doi: 10.1097/JTO.0b013e31824cc334 [9] Kim Y,Hammerman P S,Kim J,et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients[J]. J Clin Oncol,2014,32(2):121-128. doi: 10.1200/JCO.2013.50.8556 [10] Niu Z,Jin R,Zhang Y,et al. Signaling pathways and targeted therapies in lung squamous cell carcinoma: mechanisms and clinical trials[J]. Signal Transduct Target Ther,2022,7(1):353. doi: 10.1038/s41392-022-01200-x [11] Shi Q,Ruan J,Yang Y C,et al. rs66651343 and rs12909095 confer lung cancer risk by regulating CCNDBP1 expression[J]. PLoS One,2023,18(4):e0284347. doi: 10.1371/journal.pone.0284347 [12] Anjum J,Mitra S,Das R,et al. A renewed concept on the MAPK signaling pathway in cancers: polyphenols as a choice of therapeutics[J]. Pharmacol Res,2022,184(2022):106398. [13] Qi M, Elion E A. MAP kinase pathways [J]. J Cell Sci, 2005, 118(Pt 16): 3569-3572. [14] Morrison D K. MAP kinase pathways[J]. Cold Spring Harb Perspect Biol,2012,4(11):a011254. [15] Keshet Y, Seger R. The MAP kinase signaling cascades: A system of hundreds of components regulates a diverse array of physiological functions[M]. //Seger R. MAP kinase signaling protocols. Second Edition. Totowa, NJ: Humana Press, 2010: 3-38. [16] Sinkala M,Nkhoma P,Mulder N,et al. Integrated molecular characterisation of the MAPK pathways in human cancers reveals pharmacologically vulnerable mutations and gene dependencies[J]. Commun Biol,2021,4(1):9. doi: 10.1038/s42003-020-01552-6 [17] Brennecke J,Stark A,Russell RB,et al. Principles of microRNA-target recognition[J]. PLoS Biol,2005,3(3):e85. doi: 10.1371/journal.pbio.0030085 [18] Liu C J,Fu X,Xia M,et al. miRNASNP-v3: a comprehensive database for SNPs and disease-related variations in miRNAs and miRNA targets[J]. Nucleic Acids Research,2021,49(D1):D1276-D1281. doi: 10.1093/nar/gkaa783 [19] Wu W,Wu L,Zhu M,et al. miRNA mediated noise making of 3'UTR mutations in cancer[J]. Genes (Basel),2018,9(11):545. doi: 10.3390/genes9110545 [20] 中华医学会,中华医学会肿瘤学分会,中华医学会杂志社. 中华医学会肺癌临床诊疗指南(2018版)[J]. 中华肿瘤杂志,2018,40(12):30. [21] 赫捷,李霓,陈万青,等. 中国肺癌筛查与早诊早治指南(2021,北京)[J]. 中国肿瘤,2021,30(02):81-111. [22] 黄鼎智, 李琳, 李旭, 等.老年晚期肺癌内科治疗中国专家共识(2022版)[J].中国肺癌杂志, 2022, 25(06): 363-384. [23] Yang J,Yan Z,Wang Y,et al. Association study of relationships of polymorphisms in the miR-21,miR-26b,miR-221/222 and miR-126 genes with cervical intraepithelial neoplasia and cervical cancer[J]. BMC Cancer,2021,21(1):997. doi: 10.1186/s12885-021-08743-2 [24] Shi Y Y,He L. SHEsis,a powerful software platform for analyses of linkage disequilibrium,haplotype construction,and genetic association at polymorphism loci[J]. Cell Res,2005,15(2):97-98. doi: 10.1038/sj.cr.7290272 [25] Lee J,Son M J,Son C Y,et al. Genetic Variation and Autism: A Field Synopsis and Systematic Meta-Analysis[J]. Brain Sci,2020,10(10):692. doi: 10.3390/brainsci10100692 [26] Lake D,Corrêa S A,Müller J. Negative feedback regulation of the ERK1/2 MAPK pathway[J]. Cell Mol Life Sci,2016,73(23):4397-413. doi: 10.1007/s00018-016-2297-8 [27] Balmanno K,Cook S J. Tumour cell survival signalling by the ERK1/2 pathway[J]. Cell Death & Differentiation,2009,16(3):368-377. [28] Yan Z,Ohuchida K,Fei S,et al. Inhibition of ERK1/2 in cancer-associated pancreatic stellate cells suppresses cancer-stromal interaction and metastasis[J]. BioMed Central,2019,38(1):221. [29] Marampon F,Ciccarelli C,Zani B M. Biological rationale for targeting MEK/ERK pathways in anti-cancer therapy and to potentiate tumour responses to radiation[J]. Int J Mol Sci,2019,20(10):2530. doi: 10.3390/ijms20102530 [30] Zhou B,Lin W,Long Y,et al. Notch signaling pathway: architecture,disease,and therapeutics[J]. Signal Transduct Target Ther,2022,7(1):95. doi: 10.1038/s41392-022-00934-y [31] Pino M S,Chung D C. The chromosomal instability pathway in colon cancer[J]. Gastroenterology,2010,138(6):2059-2072. doi: 10.1053/j.gastro.2009.12.065 [32] Müller M F,Ibrahim A E,Arends M J. Molecular pathological classification of colorectal cancer[J]. Virchows Arch,2016,469(2):125-134. doi: 10.1007/s00428-016-1956-3 [33] Ding L,Getz G,Wheeler D A,et al. Somatic mutations affect key pathways in lung adenocarcinoma[J]. Nature,2008,455(7216):1069-1075. doi: 10.1038/nature07423 [34] Hill M,Tran N. miRNA interplay: mechanisms and consequences in cancer[J]. Dis Model Mech,2021,14(4):dmm047662. doi: 10.1242/dmm.047662 [35] Zhu Z,Zhang F,Hu H,et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets[J]. Nat Genet,2016,48(5):481-487. doi: 10.1038/ng.3538 [36] Fabian M R,Sonenberg N. The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC[J]. Nat Struct Mol Biol,2012,19(6):586-593. doi: 10.1038/nsmb.2296 [37] Chan J J,Tabatabaeian H,Tay Y. 3'UTR heterogeneity and cancer progression[J]. Trends Cell Biol,2023,33(7):568-582. doi: 10.1016/j.tcb.2022.10.001 [38] Sturgill T W,Ray L B,Erikson E,et al. Insulin-stimulated MAP-2 kinase phosphorylates and activates ribosomal protein S6 kinase II[J]. Nature,1988,334(6184):715-718. doi: 10.1038/334715a0 [39] AACR Project GENIE Consortium. AACR project GENIE: powering precision medicine through an international consortium[J]. Cancer Discov,2017,7(8):818-831. doi: 10.1158/2159-8290.CD-17-0151 [40] Rubio K,Romero-Olmedo A J,Sarvari P,et al. Non-canonical integrin signaling activates EGFR and RAS-MAPK-ERK signaling in small cell lung cancer[J]. Theranostics,2023,13(8):2384-2407. doi: 10.7150/thno.79493 [41] Wang Y,Guo Z,Tian Y,et al. MAPK1 promotes the metastasis and invasion of gastric cancer as a bidirectional transcription factor[J]. BMC Cancer,2023,23(1):959. doi: 10.1186/s12885-023-11480-3 [42] Zhu L,Yang S,Wang J. miR-217 inhibits the migration and invasion of HeLa cells through modulating MAPK1[J]. Int J Mol Med,2019,44(5):1824-1832. [43] Gagliardi M,Pitner M K,Park J,et al. Differential functions of ERK1 and ERK2 in lung metastasis processes in triple-negative breast cancer[J]. Sci Rep,2020,10(1):8537. doi: 10.1038/s41598-020-65250-3 [44] Guo N,Zhang N,Yan L,et al. Correlation between genetic polymorphisms within the MAPK1/HIF-1/HO-1 signaling pathway and risk or prognosis of perimenopausal coronary artery disease[J]. Clin Cardiol,2017,40(8):597-604. doi: 10.1002/clc.22708 [45] Insodaite R,Smalinskiene A,Liutkevicius V,et al. Associations of polymorphisms localized in the 3'UTR regions of the KRAS,NRAS,MAPK1 genes with laryngeal squamous cell carcinoma[J]. Genes (Basel),2021,12(11):1679. doi: 10.3390/genes12111679 [46] Hirsch F R,Scagliotti G V,Mulshine J L,et al. Lung cancer: current therapies and new targeted treatments[J]. Lancet,2017,389(10066):299-311. doi: 10.1016/S0140-6736(16)30958-8 [47] Kulasingam V,Diamandis E P. Strategies for discovering novel cancer biomarkers through utilization of emerging technologies[J]. Nat Clin Pract Oncol,2008,5(10):588-599. doi: 10.1038/ncponc1187 [48] Chen W,Zheng R,Baade P D,et al. Cancer statistics in China,2015[J]. CA Cancer J Clin,2016,66(2):115-132. doi: 10.3322/caac.21338 -




 下载:
下载: